Mutation: Difference between revisions
imported>John J. Dennehy No edit summary |
imported>John J. Dennehy No edit summary |
||
| Line 1: | Line 1: | ||
In biology, mutations are changes | == Overview == | ||
In [[biology]], mutations are changes in the [[DNA]] or [[RNA]] sequences of organisms caused by copying errors during [[DNA replication]] or by damage from exposure to [[radiation]], chemicals, or [[virus]]es. Mutations, ultimately, are the only source of [[heritable variation]] on which [[natural selection]] and other [[evolution]]ary processes acts. Without mutation, there are no new [[gene]]s, no new [[allele]]s and, eventually, no evolutionary change. | |||
Mutations create variations in the gene pool | Mutations create variations in the [[gene pool]]. The basis of evolutionary change is the removal of unfavorable mutations from and the retention of favorable mutations in the gene pool by natural selection. Neutral mutations are defined as mutations that do not affect organismal [[fitness]]; they are, by definition, invisible to natural selection. | ||
== Contents == | |||
1 Classification | |||
o 1.1 By effect on structure | o 1.1 By effect on structure | ||
o 1.2 By effect on function | o 1.2 By effect on function | ||
| Line 12: | Line 15: | ||
o 1.5 Causes of mutation | o 1.5 Causes of mutation | ||
o 1.6 Nomenclature | o 1.6 Nomenclature | ||
2 Harmful mutations | |||
3 References | |||
o 3.1 Online books | |||
4 External links | |||
== Classification == | |||
By effect on structure | |||
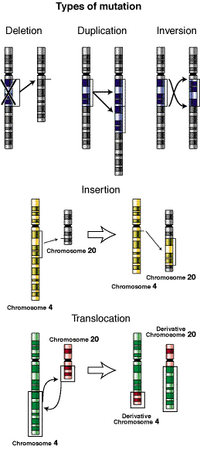
[[Image:Types-of-mutation.png|thumb|200 px|Illustrations of five types of chromosomal mutations.]] | |||
Illustrations of five types of chromosomal mutations. | |||
DNA sequences can be altered in a number of ways and have varying effects on health depending on where they occur and whether they alter the function of essential [[protein]]s or [[regulatory elements]]. Structurally, mutations can be classified as: | |||
*Small-scale mutations, such as affecting a small gene is one or a few nucleotides, including: | |||
o [[Point mutations]], often caused by chemicals or DNA replication errors, exchange a single [[nucleotide]] for another. Most common is the transition that exchanges a [[purine]] for a purine (A ↔ G) or a [[pyrimidine]] for a pyrimidine, (C ↔ T). A transition can be caused by nitrous acid, base mispairing, or mutagenic base analogs such as 5-bromo-2-deoxyuridine (BrdU). Less common is a [[transversion]], which exchanges a purine for a pyrimidine or a pyrimidine for a purine (C/T ↔ A/G). A point mutation can be reversed by another point mutation, in which the nucleotide is changed back to its original state (true reversion) or by second-site reversion (a complementary mutation elsewhere that results in regained gene functionality). These changes are classified as [[transitions]] or transversions. An example of a transversion is [[adenine]] (A) being converted into a [[cytosine]] (C). There are also many other examples that can be found. Point mutations that occur within the protein coding region of a gene may be classified into three kinds, depending upon what the erroneous [[codon]] codes for: | |||
+ [[Silent mutations]]: which code for the same [[amino acid]]. | |||
+ [[Missense mutations]]: which code for a different amino acid. | |||
+ [[Nonsense mutations]]: which code for a stop codon and can truncate a protein. | |||
o Insertions add one or more extra nucleotides into the DNA and are usually caused by [[transposable element]]s or errors during replication of repeating elements (e.g., AT repeats). Insertions in the coding region of a gene may alter splicing of the [[mRNA]] (splice site mutation), or cause a shift in the [[reading frame]] (frameshift), both of which can significantly alter the gene product. Insertions can be reverted by excision of the transposable element. | |||
o Deletions remove one or more nucleotides from the DNA. Like insertions, these mutations can alter the reading frame of the gene. They are irreversible. | |||
* Large-scale mutations in chromosomal structure, including: | |||
o Amplifications (or gene duplications) leading to multiple copies of all chromosomal regions, increasing the dosage of the genes located within them. | |||
o Deletions of large chromosomal regions, leading to loss of the genes within those regions. | |||
o Mutations whose effect is to juxtapose previously separate pieces of DNA, potentially bringing together separate genes to form functionally distinct fusion genes (e.g. bcr-abl). These include: | |||
+ Chromosomal translocations: interchange of genetic parts from non[[homologous]] chromosomes. | |||
+ Interstitial deletions: an intra-chromosomal deletion that removes a segment of DNA from a single chromosome, thereby apposing previously distant genes. For example, cells isolated from a [[human astrocytoma]], a type of brain [[tumor]], were found to have a chromosomal deletion removing sequences between the "fused in glioblastoma" (fig) gene and the receptor tyrosine kinase "ros", producing a fusion protein (FIG-ROS). The abnormal FIG-ROS fusion protein has constitutively active kinase activity that causes oncogenic transformation (a transformation from normal cells to cancer cells). (citation) | |||
+ Chromosomal inversions: reversing the orientation of a chromosomal segment. | |||
o Loss of [[heterozygosity]]: loss of one allele, either by a deletion or [[recombination]] event, in an organism that previously had two different [[allele]]s. | |||
== By effect on function == | |||
* Loss-of-function mutations are the result of gene product having less or no function. When the allele has a complete loss of function (null allele) it is often called an [[amorphic]] mutation. [[Phenotype]]s associated with such mutations are most often [[recessive]]. Exceptions are when the organism is [[haploid]], or when the reduced dosage of a normal gene product is not enough for a normal phenotype (this is called haploinsufficiency). | |||
* Gain-of-function mutations change the gene product such that it gains a new and abnormal function. These mutations usually have [[dominant]] phenotypes. Often called a [[neomorphic]] mutation. | |||
* Dominant negative mutations (also called antimorphic mutations) have an altered gene product that acts antagonistically to the wild-type allele. These mutations usually result in an altered molecular function (often inactive) and are | * Dominant negative mutations (also called [[antimorphic]] mutations) have an altered gene product that acts antagonistically to the wild-type allele. These mutations usually result in an altered molecular function (often inactive) and are characterized by a dominant or semi-dominant phenotype. In humans, [[Marfan's syndrome]] is an example of a dominant negative mutation occurring in an autosomal dominant disease. In this condition, the defective [[glycoprotein]] product of the fibrillin gene (FBN1) antagonizes the product of the normal allele. | ||
* Lethal mutations are mutations that lead to a phenotype incapable of effective reproduction. | |||
== By aspect of phenotype affected == | |||
* [[Morphological]] mutations usually affect the outward appearance of an individual. Mutations can change the height of a plant or change it from smooth to rough seeds. | |||
* [[Biochemical]] mutations result in lesions stopping the [[enzymatic pathway]]. Often, morphological mutants are the direct result of a mutation due to the enzymatic pathway. | |||
== Special classes == | |||
* Conditional mutation is a mutation that has wild-type (or less severe) phenotype under certain "permissive" [[environment]]al conditions and a mutant phenotype under certain "restrictive" conditions. For example, a temperature-sensitive mutation can cause cell death at high temperature (restrictive condition), but might have no deleterious consequences at a lower temperature (permissive condition). | |||
== Causes of mutation == | |||
Two classes of mutations are spontaneous mutations (molecular decay) and induced mutations caused by | Two classes of mutations are spontaneous mutations (molecular decay) and induced mutations caused by [[mutagen]]s. | ||
Spontaneous mutations on the molecular level include: | Spontaneous mutations on the molecular level include: | ||
* [[Tautomerism]] - A base is changed by the repositioning of a [[hydrogen]] [[atom]]. | |||
* [[Depurination]] - Loss of a purine base (A or G). | |||
* [[Deamination]] - Changes a normal base to an atypical base; C → U, (which can be corrected by DNA repair mechanisms), or spontaneous deamination of 5-methycytosine (irreparable), or A → HX (hypoxanthine). | |||
* Transition - A purine changes to another purine, or a pyrimidine to a pyrimidine. | |||
* Transversion - A purine becomes a pyrimidine, or vice versa. | |||
Induced mutations on the molecular level can be caused by: | Induced mutations on the molecular level can be caused by: | ||
* Chemicals | |||
o Nitrosoguanidine (NTG) | |||
o Hydroxyamine NH3OH | |||
o Base analogs (e.g. BrdU) | |||
o Simple chemicals (e.g. acids) | |||
o Alkylating agents (e.g. N-ethyl-N-nitrosourea (ENU)) These agents can mutate both replicating and non-replicating DNA. In contrast, a base analog can only mutate the DNA when the analog is incorporated in replicating the DNA. Each of these classes of chemical mutagens has certain effects that then lead to transitions, transversions, or deletions. | |||
o Methylating agents (e.g. ethyl methanesulfonate (EMS)) | |||
o Polycyclic hydrocarbons (e.g. benzopyrenes found in internal combustion engine exhaust) | |||
o DNA intercalating agents (e.g. ethidium bromide) | |||
o DNA crosslinker (e.g. platinum) | |||
o Oxidative damage caused by oxygen(O)] radicals | |||
* Radiation | |||
o Ultraviolet radiation (nonionizing radiation) - excites electrons to a higher energy level. DNA absorbs one form, [[ultraviolet light]]. Two nucleotide bases in DNA - cytosine and thymine - are most vulnerable to excitation that can change base-pairing properties. UV light can induce adjacent thymine bases in a DNA strand to pair with each other, as a bulky [[dimer]]. | |||
o Ionizing radiation | |||
DNA has so-called hotspots, where mutations occur up to 100 times more frequently than the normal mutation rate. A hotspot can be at an unusual base, e.g., 5-methylcytosine. | DNA has so-called [[hotspots]], where mutations occur up to 100 times more frequently than the normal [[mutation rate]]. A hotspot can be at an unusual base, e.g., 5-methylcytosine. | ||
Mutation rates also vary across species. Evolutionary biologists have theorized that higher mutation rates are beneficial in some situations | Mutation rates also vary across [[species]]. Evolutionary biologists have theorized that higher mutation rates are beneficial in some situations because they allow organisms to evolve and therefore [[adapt]] more quickly to their environments. For example, repeated exposure of [[bacteria]] to [[antibiotic]]s, and selection of resistant mutants, can result in the selection of bacteria that have a much higher mutation rate than the original population (mutator strains). (citation) | ||
== Nomenclature == | |||
Nomenclature of mutations specify the type of mutation and base or amino acid changes. | Nomenclature of mutations specify the type of mutation and base or amino acid changes. | ||
* Amino acid substitution - (e.g. D111E) The first letter is the one letter code of the wildtype amino acid, the number is the position of the amino acid from the N terminus and the second letter is the one letter code of the amino acid present in the mutation. If the second letter is 'X', any amino acid may replace the wildtype. | |||
* Amino acid deletion - (e.g. ΔF508) The Greek symbol Δ or 'delta' indicates a deletion. The letter refers to the amino acid present in the wildtype and the number is the position from the N terminus of the amino acid were it to be present as in the wildtype. | |||
== Harmful mutations == | |||
Changes in DNA caused by mutation can cause errors in protein sequence, creating partially or completely non-functional proteins. To function correctly, each cell depends on thousands of proteins to function in the right places at the right times. When a mutation alters a protein that plays a critical role in the body, a medical condition can result. A condition caused by mutations in one or more genes is called a genetic disorder. However, only a small percentage of mutations cause genetic disorders; most have no impact on health. For example, some mutations alter a gene's DNA base sequence but don’t change the function of the protein made by the gene. | Changes in DNA caused by mutation can cause errors in protein sequence, creating partially or completely non-functional proteins. To function correctly, each cell depends on thousands of proteins to function in the right places at the right times. When a mutation alters a protein that plays a critical role in the body, a medical condition can result. A condition caused by mutations in one or more genes is called a genetic disorder. However, only a small percentage of mutations cause genetic disorders; most have no impact on health. For example, some mutations alter a gene's DNA base sequence but don’t change the function of the protein made by the gene. | ||
If a mutation is present in a germ cell, it can give rise to offspring that carries the mutation in all of its cells. This is the case in hereditary | If a mutation is present in a [[germ cell]], it can give rise to offspring that carries the mutation in all of its cells. This is the case in [[hereditary disease]]s. On the other hand, a mutation can occur in a [[somatic cell]] of an organism. Such mutations will be present in all descendants of this cell, and certain mutations can cause the cell to become [[malignant]], and thus cause [[cancer]]. | ||
Often, gene mutations that could cause a genetic disorder are repaired by the DNA repair system of the cell. Each cell has a number of pathways through which enzymes recognize and repair mistakes in DNA. Because DNA can be damaged or mutated in many ways, the process of DNA repair is an important way in which the body protects itself from disease. | Often, gene mutations that could cause a genetic disorder are repaired by the DNA repair system of the cell. Each cell has a number of pathways through which enzymes recognize and repair mistakes in DNA. Because DNA can be damaged or mutated in many ways, the process of DNA repair is an important way in which the body protects itself from disease. | ||
== References == | |||
* Leroi A. 2003. Mutants: On the form, varieties & errors of the human body. 1:16-17. Harper Collins 2003 | |||
* Maki H. 2002. Origins of spontaneous mutations: specificity and directionality of base-substitution, frameshift, and sequence-substitution mutageneses. Annual Review of Genetics 36:279-303. | |||
* Taggart R. Starr C. Biology The Unity and Diversity of Life: Mutated Genes and Their Protein Products. 14.4:227. Thompson Brooks/Cole 2006. | |||
[ | == Online books == | ||
* Chapter 7, The Molecular Basis of Mutation in Modern Genetic Analysis by Anthony J. F. Griffiths, William M. Gelbart, Jeffrey H. Miller and Richard C. Lewontin (1999) published by W. H. Freeman and Company ISBN 0-7167-3597-0. | |||
[http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Search&db=books&doptcmdl=GenBookHL&term=mutation+AND+mga%5Bbook%5D+AND+110363%5Buid%5D&rid=mga.section.996 The Molecular Basis of Mutation] | |||
* Chapter 9, Instability of the human genome: mutation and DNA repair in Human Molecular Genetics 2 by Tom Strachan and Andrew P. Read (1999) published by John Wiley & Sons, Inc. [http://www.ncbi.nlm.nih.gov/books/bv.fcgi?rid=hmg.section.1050 Instability of the human genome: mutation and DNA repair] | |||
[ | * Genes and Disease from the National Library of Medicine provides descriptions of mutations that cause human diseases. For example, a common mutation associated with Huntington disease is an increased number of copies of repeated CGA triplets in the Huntingtin gene.[http://www.ncbi.nlm.nih.gov/books/bv.fcgi?rid=gnd.preface.91 Genes and Disease] | ||
* GeneReviews by Roberta A. Pagon, Editor-in-chief is made available by the University of Washington and contains peer-reviewed descriptions of heritable diseases written by experts. For example, BRCA1 and BRCA2 Hereditary Breast/Ovarian Cancer describes mutations in BRCA1 and BRCA2 that are associated with predispositions to cancer.[http://www.ncbi.nlm.nih.gov/books/bv.fcgi?rid=gene GeneReviews] | |||
Revision as of 21:07, 19 June 2007
Overview
In biology, mutations are changes in the DNA or RNA sequences of organisms caused by copying errors during DNA replication or by damage from exposure to radiation, chemicals, or viruses. Mutations, ultimately, are the only source of heritable variation on which natural selection and other evolutionary processes acts. Without mutation, there are no new genes, no new alleles and, eventually, no evolutionary change.
Mutations create variations in the gene pool. The basis of evolutionary change is the removal of unfavorable mutations from and the retention of favorable mutations in the gene pool by natural selection. Neutral mutations are defined as mutations that do not affect organismal fitness; they are, by definition, invisible to natural selection.
Contents
1 Classification
o 1.1 By effect on structure
o 1.2 By effect on function
o 1.3 By aspect of phenotype affected
o 1.4 Special classes
o 1.5 Causes of mutation
o 1.6 Nomenclature
2 Harmful mutations 3 References o 3.1 Online books 4 External links
Classification
By effect on structure
DNA sequences can be altered in a number of ways and have varying effects on health depending on where they occur and whether they alter the function of essential proteins or regulatory elements. Structurally, mutations can be classified as:
- Small-scale mutations, such as affecting a small gene is one or a few nucleotides, including:
o Point mutations, often caused by chemicals or DNA replication errors, exchange a single nucleotide for another. Most common is the transition that exchanges a purine for a purine (A ↔ G) or a pyrimidine for a pyrimidine, (C ↔ T). A transition can be caused by nitrous acid, base mispairing, or mutagenic base analogs such as 5-bromo-2-deoxyuridine (BrdU). Less common is a transversion, which exchanges a purine for a pyrimidine or a pyrimidine for a purine (C/T ↔ A/G). A point mutation can be reversed by another point mutation, in which the nucleotide is changed back to its original state (true reversion) or by second-site reversion (a complementary mutation elsewhere that results in regained gene functionality). These changes are classified as transitions or transversions. An example of a transversion is adenine (A) being converted into a cytosine (C). There are also many other examples that can be found. Point mutations that occur within the protein coding region of a gene may be classified into three kinds, depending upon what the erroneous codon codes for: + Silent mutations: which code for the same amino acid. + Missense mutations: which code for a different amino acid. + Nonsense mutations: which code for a stop codon and can truncate a protein. o Insertions add one or more extra nucleotides into the DNA and are usually caused by transposable elements or errors during replication of repeating elements (e.g., AT repeats). Insertions in the coding region of a gene may alter splicing of the mRNA (splice site mutation), or cause a shift in the reading frame (frameshift), both of which can significantly alter the gene product. Insertions can be reverted by excision of the transposable element. o Deletions remove one or more nucleotides from the DNA. Like insertions, these mutations can alter the reading frame of the gene. They are irreversible.
- Large-scale mutations in chromosomal structure, including:
o Amplifications (or gene duplications) leading to multiple copies of all chromosomal regions, increasing the dosage of the genes located within them. o Deletions of large chromosomal regions, leading to loss of the genes within those regions. o Mutations whose effect is to juxtapose previously separate pieces of DNA, potentially bringing together separate genes to form functionally distinct fusion genes (e.g. bcr-abl). These include: + Chromosomal translocations: interchange of genetic parts from nonhomologous chromosomes. + Interstitial deletions: an intra-chromosomal deletion that removes a segment of DNA from a single chromosome, thereby apposing previously distant genes. For example, cells isolated from a human astrocytoma, a type of brain tumor, were found to have a chromosomal deletion removing sequences between the "fused in glioblastoma" (fig) gene and the receptor tyrosine kinase "ros", producing a fusion protein (FIG-ROS). The abnormal FIG-ROS fusion protein has constitutively active kinase activity that causes oncogenic transformation (a transformation from normal cells to cancer cells). (citation) + Chromosomal inversions: reversing the orientation of a chromosomal segment. o Loss of heterozygosity: loss of one allele, either by a deletion or recombination event, in an organism that previously had two different alleles.
By effect on function
- Loss-of-function mutations are the result of gene product having less or no function. When the allele has a complete loss of function (null allele) it is often called an amorphic mutation. Phenotypes associated with such mutations are most often recessive. Exceptions are when the organism is haploid, or when the reduced dosage of a normal gene product is not enough for a normal phenotype (this is called haploinsufficiency).
- Gain-of-function mutations change the gene product such that it gains a new and abnormal function. These mutations usually have dominant phenotypes. Often called a neomorphic mutation.
* Dominant negative mutations (also called antimorphic mutations) have an altered gene product that acts antagonistically to the wild-type allele. These mutations usually result in an altered molecular function (often inactive) and are characterized by a dominant or semi-dominant phenotype. In humans, Marfan's syndrome is an example of a dominant negative mutation occurring in an autosomal dominant disease. In this condition, the defective glycoprotein product of the fibrillin gene (FBN1) antagonizes the product of the normal allele.
- Lethal mutations are mutations that lead to a phenotype incapable of effective reproduction.
By aspect of phenotype affected
- Morphological mutations usually affect the outward appearance of an individual. Mutations can change the height of a plant or change it from smooth to rough seeds.
- Biochemical mutations result in lesions stopping the enzymatic pathway. Often, morphological mutants are the direct result of a mutation due to the enzymatic pathway.
Special classes
- Conditional mutation is a mutation that has wild-type (or less severe) phenotype under certain "permissive" environmental conditions and a mutant phenotype under certain "restrictive" conditions. For example, a temperature-sensitive mutation can cause cell death at high temperature (restrictive condition), but might have no deleterious consequences at a lower temperature (permissive condition).
Causes of mutation
Two classes of mutations are spontaneous mutations (molecular decay) and induced mutations caused by mutagens.
Spontaneous mutations on the molecular level include:
- Tautomerism - A base is changed by the repositioning of a hydrogen atom.
- Depurination - Loss of a purine base (A or G).
- Deamination - Changes a normal base to an atypical base; C → U, (which can be corrected by DNA repair mechanisms), or spontaneous deamination of 5-methycytosine (irreparable), or A → HX (hypoxanthine).
- Transition - A purine changes to another purine, or a pyrimidine to a pyrimidine.
- Transversion - A purine becomes a pyrimidine, or vice versa.
Induced mutations on the molecular level can be caused by:
- Chemicals
o Nitrosoguanidine (NTG) o Hydroxyamine NH3OH o Base analogs (e.g. BrdU) o Simple chemicals (e.g. acids) o Alkylating agents (e.g. N-ethyl-N-nitrosourea (ENU)) These agents can mutate both replicating and non-replicating DNA. In contrast, a base analog can only mutate the DNA when the analog is incorporated in replicating the DNA. Each of these classes of chemical mutagens has certain effects that then lead to transitions, transversions, or deletions. o Methylating agents (e.g. ethyl methanesulfonate (EMS)) o Polycyclic hydrocarbons (e.g. benzopyrenes found in internal combustion engine exhaust) o DNA intercalating agents (e.g. ethidium bromide) o DNA crosslinker (e.g. platinum) o Oxidative damage caused by oxygen(O)] radicals
- Radiation
o Ultraviolet radiation (nonionizing radiation) - excites electrons to a higher energy level. DNA absorbs one form, ultraviolet light. Two nucleotide bases in DNA - cytosine and thymine - are most vulnerable to excitation that can change base-pairing properties. UV light can induce adjacent thymine bases in a DNA strand to pair with each other, as a bulky dimer. o Ionizing radiation
DNA has so-called hotspots, where mutations occur up to 100 times more frequently than the normal mutation rate. A hotspot can be at an unusual base, e.g., 5-methylcytosine.
Mutation rates also vary across species. Evolutionary biologists have theorized that higher mutation rates are beneficial in some situations because they allow organisms to evolve and therefore adapt more quickly to their environments. For example, repeated exposure of bacteria to antibiotics, and selection of resistant mutants, can result in the selection of bacteria that have a much higher mutation rate than the original population (mutator strains). (citation)
Nomenclature
Nomenclature of mutations specify the type of mutation and base or amino acid changes.
- Amino acid substitution - (e.g. D111E) The first letter is the one letter code of the wildtype amino acid, the number is the position of the amino acid from the N terminus and the second letter is the one letter code of the amino acid present in the mutation. If the second letter is 'X', any amino acid may replace the wildtype.
- Amino acid deletion - (e.g. ΔF508) The Greek symbol Δ or 'delta' indicates a deletion. The letter refers to the amino acid present in the wildtype and the number is the position from the N terminus of the amino acid were it to be present as in the wildtype.
Harmful mutations
Changes in DNA caused by mutation can cause errors in protein sequence, creating partially or completely non-functional proteins. To function correctly, each cell depends on thousands of proteins to function in the right places at the right times. When a mutation alters a protein that plays a critical role in the body, a medical condition can result. A condition caused by mutations in one or more genes is called a genetic disorder. However, only a small percentage of mutations cause genetic disorders; most have no impact on health. For example, some mutations alter a gene's DNA base sequence but don’t change the function of the protein made by the gene.
If a mutation is present in a germ cell, it can give rise to offspring that carries the mutation in all of its cells. This is the case in hereditary diseases. On the other hand, a mutation can occur in a somatic cell of an organism. Such mutations will be present in all descendants of this cell, and certain mutations can cause the cell to become malignant, and thus cause cancer.
Often, gene mutations that could cause a genetic disorder are repaired by the DNA repair system of the cell. Each cell has a number of pathways through which enzymes recognize and repair mistakes in DNA. Because DNA can be damaged or mutated in many ways, the process of DNA repair is an important way in which the body protects itself from disease.
References
- Leroi A. 2003. Mutants: On the form, varieties & errors of the human body. 1:16-17. Harper Collins 2003
- Maki H. 2002. Origins of spontaneous mutations: specificity and directionality of base-substitution, frameshift, and sequence-substitution mutageneses. Annual Review of Genetics 36:279-303.
- Taggart R. Starr C. Biology The Unity and Diversity of Life: Mutated Genes and Their Protein Products. 14.4:227. Thompson Brooks/Cole 2006.
Online books
- Chapter 7, The Molecular Basis of Mutation in Modern Genetic Analysis by Anthony J. F. Griffiths, William M. Gelbart, Jeffrey H. Miller and Richard C. Lewontin (1999) published by W. H. Freeman and Company ISBN 0-7167-3597-0.
The Molecular Basis of Mutation
- Chapter 9, Instability of the human genome: mutation and DNA repair in Human Molecular Genetics 2 by Tom Strachan and Andrew P. Read (1999) published by John Wiley & Sons, Inc. Instability of the human genome: mutation and DNA repair
- Genes and Disease from the National Library of Medicine provides descriptions of mutations that cause human diseases. For example, a common mutation associated with Huntington disease is an increased number of copies of repeated CGA triplets in the Huntingtin gene.Genes and Disease
- GeneReviews by Roberta A. Pagon, Editor-in-chief is made available by the University of Washington and contains peer-reviewed descriptions of heritable diseases written by experts. For example, BRCA1 and BRCA2 Hereditary Breast/Ovarian Cancer describes mutations in BRCA1 and BRCA2 that are associated with predispositions to cancer.GeneReviews